One of the dominating features of the economy over the last fifty years has been Moore’s law, which has led to exponential growth in computing power and exponential drops in its costs. This period is now coming to an end. This doesn’t mean that technological progress in computing will stop dead, nor that innovation in ICT comes to an end, but it is a pivotal change, and I’m surprised that we’re not seeing more discussion of its economic implications.
This reflects, perhaps, the degree to which some economists seem to be both ill-informed and incurious about the material and technical basis of technological innovation (for a very prominent example, see my review of Robert Gordon’s recent widely read book, The Rise and Fall of American Growth). On the other hand, boosters of the idea of accelerating change are happy to accept it as axiomatic that these technological advances will always continue at the same, or faster rates. Of course, the future is deeply uncertain, and I am not going to attempt to make many predictions. But here’s my attempt to identify some of the issues
How we got here
The era of Moore’s law began with the invention in 1959 of the integrated circuit. Transistors are the basic unit of electronic unit, and in an integrated circuit many transistors could be incorporated on a single component to make a functional device. As the technology for making integrated circuits rapidly improved, Gordon Moore, in 1965 predicted that the number of transistors on a single silicon chip would double every year (the doubling time was later revised to 18 months, but in this form the “law” has well described the products of the semiconductor industry ever since).
The full potential of integrated circuits was realised when, in effect, a complete computer was built on a single chip of silicon – a microprocessor. The first microprocessor was made in 1970, to serve as the flight control computer for the F14 Tomcat. Shortly afterwards a civilian microprocessor was released by Intel – the 4004. This was followed in 1974 by the Intel 8080 and its competitors, which were the devices that launched the personal computer revolution.
The Intel 8080 had transistors with a minimum feature size of 6 µm. Moore’s law was driven by a steady reduction in this feature size – by 2000, Intel’s Pentium 4’s transistors were more than 30 times smaller. This drove the huge increase in computer power between the two chips in two ways. Obviously, more transistors gives you more logic gates, and more is better. Less obviously, another regularity known as Dennard scaling states that as transistor dimensions are shrunk, each transistor operates faster and uses less power. The combination of Moore’s law and Dennard scaling was what led to the golden age of microprocessors, from the mid-1990’s, where every two years a new generation of technology would be introduced, each one giving computers that were cheaper and faster than the last.
This golden age began to break down around 2004. Transistors were still shrinking, but the first physical limit was encountered. Further increases in speed became impossible to sustain, because the processors simply ran too hot. To get round this, a new strategy was introduced – the introduction of multiple cores. The transistors weren’t getting much faster, but more computer power came from having more of them – at the cost of some software complexity. This marked a break in the curve of improvement of computer power with time, as shown in the figure below.
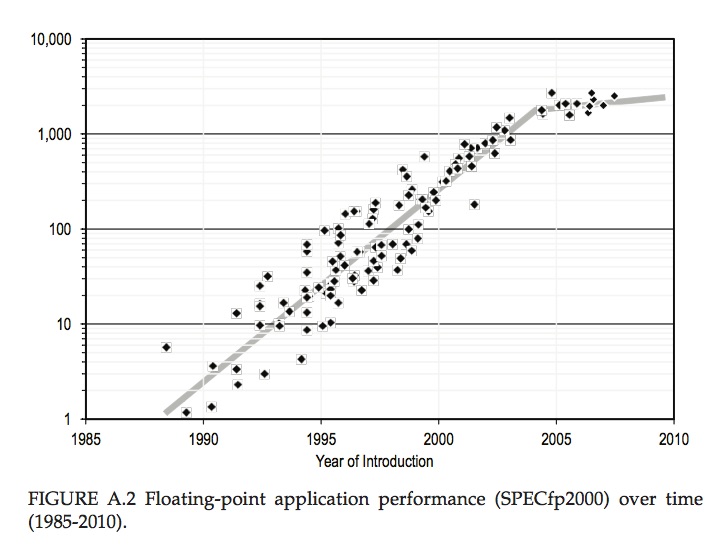
Computer performance trends as measured by the SPECfp2000 standard for floating point performance, normalised to a typical 1985 value. This shows an exponential growth in computer power from 1985 to 2004 at a compound annual rate exceeding 50%, and slower growth between 2004 and 2010. From “The Future of Computing Performance: Game Over or Next Level?”, National Academies Press 2011″
In this period, transistor dimensions were still shrinking, even if they weren’t becoming faster, and the cost per transistor was still going down. But as dimensions shrunk to tens of nanometers, chip designers ran out of space, and further increases in density were only possible by moving into the third dimension. The “FinFET” design, introduced in 2011 essentially stood the transistors on their side. At this point the reduction in cost per transistor began to level off, and since then the development cycle has begun to slow, with Intel announcing a move from a two year cycle to one of three years.
The cost of sustaining Moore’s law can be measured in diminishing returns from R&D efforts (estimated by Bloom et al as a roughly 8-fold increase in research effort, measured as R&D expenditure deflated by researcher salaries, from 1995 to 2015), and above all by rocketing capital costs.
Oligopoly concentration
The cost of the most advanced semiconductor factories (fabs) now exceeds $10 billion, with individual tools approaching $100 million. This rocketing cost of entry means that now only four companies in the world have the capacity to make semiconductor chips at the technological leading edge.
These firms are Intel (USA), Samsung (Korea), TSMC (Taiwan) and Global Foundries (USA/Singapore based, but owned by the Abu Dhabi sovereign wealth fund). Other important names in semiconductors are now “fabless” – they design chips, that are then manufactured in fabs operated by one of these four. These fabless firms include nVidia – famous for the graphical processing units that have been so important for computer games, but which are now becoming important for the high performance computing needed for AI and machine learning, and ARM (until recently UK based and owned, but recently bought by Japan’s SoftBank), designer of low power CPUs for mobile devices.
It’s not clear to me how the landscape evolves from here. Will there be further consolidation? Or in an an environment of increasing economic nationalism, will ambitious nations regard advanced semiconductor manufacture as a necessary sovereign capability, to be acquired even in the teeth of pure economic logic? Of course, I’m thinking mostly of China in this context – its government has a clearly stated policy of attaining technological leadership in advanced semiconductor manufacturing.
Cheap as chips
The flip-side of diminishing returns and slowing development cycles at the technological leading edge is that it will make sense to keep those fabs making less advanced devices in production for longer. And since so much of the cost of an IC is essentially the amortised cost of capital, once that is written off the marginal cost of making more chips in an old fab is small. So we can expect the cost of trailing edge microprocessors to fall precipitously. This provides the economic driving force for the idea of the “internet of things”. Essentially, it will be possible to provide a degree of product differentiation by introducing logic circuits into all sorts of artefacts – putting a microprocessor in every toaster, in other words.
Although there are applications where cheap embedded computing power can be very valuable, I’m not sure this is a universally good idea. There is a danger that we will accept relatively marginal benefits (the ability to switch our home lights on with our smart-phones, for example) at the price of some costs that may not be immediately obvious. There will be a general loss of transparency and robustness of everyday technologies, and the potential for some insidious potential harms, through vulnerability to hostile cyberattacks, for example. Caution is required!
Travelling without a roadmap
Another important feature of the golden age of Moore’s law and Dennard scaling was a social innovation – the International Technology Roadmap for Semiconductors. This was an important (and I think unique) device for coordinating and setting the pace for innovation across a widely dispersed industry, comprising equipment suppliers, semiconductor manufacturers, and systems integrators. The relentless cycle of improvement demanded R&D in all sorts of areas – the materials science of the semiconductors, insulators and metals and their interfaces, the chemistry of resists, the optics underlying the lithography process – and this R&D needed to be started not in time for the next upgrade, but many years in advance of when it was anticipated it would be needed. Meanwhile businesses could plan products that wouldn’t be viable with the computer power available at that time, but which could be expected in the future.
Moore’s law was a self-fulfilling prophecy, and the ITRS was the document that both predicted the future, and made sure that that future happened. I write this in the past tense, because there will be no more roadmaps. Changing industry conditions – especially the concentration of leading edge manufacturing – has brought this phase to an end, and the last International Technology Roadmap for Semiconductors was issued in 2015.
What does all this mean for the broader economy?
The impact of fifty years of exponential technological progress in computing seems obvious, yet quantifying its contribution to the economy is more difficult. In developed countries, the information and communication technology sector has itself been a major part of the economy which has demonstrated very fast productivity growth. In fact, the rapidity of technological change has itself made the measurement of economic growth more difficult, with problems arising in accounting for the huge increases in quality at a given price for personal computers, and the introduction of entirely new devices such as smartphones.
But the effects of these technological advances on the rest of the economy must surely be even larger than the direct contribution of the ICT sector. Indeed, even countries without a significant ICT industry of their own must also have benefitted from these advances. The classical theory of economic growth due to Solow can’t deal with this, as it isn’t able to handle a situation in which different areas of technology are advancing at very different rates (a situation which has been universal since at least the industrial revolution).
One attempt to deal with this was made by Oulton, who used a two-sector model to take into account the the effect of improved ICT technology in other sectors, by increasing the cost-effectiveness of ICT related capital investment in those sectors. This does allow one to make some account for the broader impact of improvements in ICT, but I still don’t think it handles the changes in relative value over time that different rates of technological improvement imply. Nonetheless, it allows one to argue for substantial contributions to economic growth from these developments.
Have we got the power?
I want to conclude with two questions for the future. I’ve already discussed the power consumption – and dissipation – of microprocessors in the context of the mid-2000’s end of Dennard scaling. Any user of a modern laptop is conscious of how much heat they generate. Aggregating the power demands of all the computing devices in the world produces a total that is a significant fraction of total energy use, and which is growing fast.
The plot below shows an estimate for the total world power consumption of ICT. This is highly approximate (and as far as the current situation goes, it looks, if anything, somewhat conservative). But it does make clear that the current trajectory is unsustainable in the context of the need to cut carbon emissions dramatically over the coming decades.
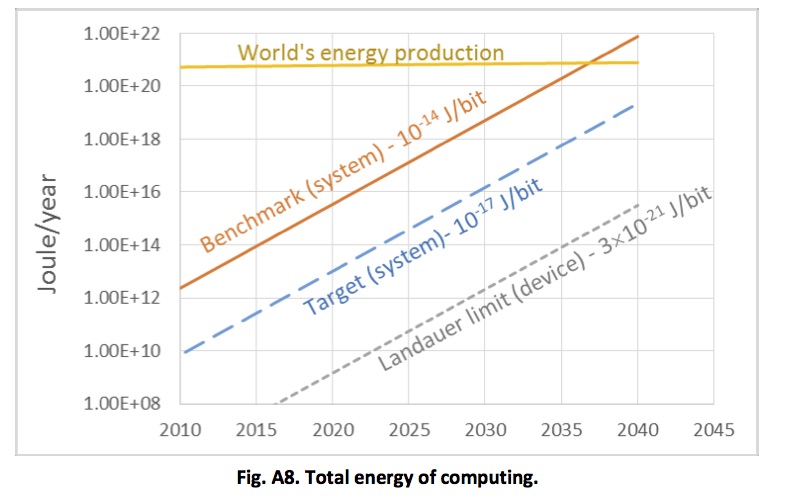
Estimated total world energy consumption for information and communication technology. From Rebooting the IT Revolution: a call to action – Semiconductor Industry Association, 2015
These rising power demands aren’t driven by more lap-tops – its the rising demands of the data centres that power the “cloud”. As smart phones became ubiquitous, we’ve seen the computing and data storage that they need move from the devices themselves, limited as they are by power consumption, to the cloud. A service like Apple’s Siri relies on technologies of natural language processing and machine learning that are much too computer intensive for the processor in the phone, and instead are run on the vast banks of microprocessors in one of Apple’s data centres.
The energy consumption of these data centres is huge and growing. By 2030, a single data centre is expected to be using 2000 MkWh per year, of which 500 MkWh is needed for cooling alone. This amounts to a power consumption of around 0.2 GW, a substantial fraction of the output of a large power station. Computer power is starting to look a little like aluminium, something that is exported from regions where electricity is cheap (and hopefully low carbon in origin). However there are limits to this concentration of computer power – the physical limit on the speed of information transfer imposed by the speed of light is significant, and the volume of information is limited by available bandwidth (especially for wireless access).
The other question is what we need that computing power for. Much of the driving force for increased computing power in recent years has been from gaming – the power needed to simulate and render realistic virtual worlds that has driven the development of powerful graphics processing units. Now it is the demands of artificial intelligence and machine learning that are straining current capacity. Truly autonomous systems, like self-driving cars, will need stupendous amounts of computer power, and presumably for true autonomy much of this computing will need to be done locally rather than in the cloud. I don’t know how big this challenge is.
Where do we go from here?
In the near term, Moore’s law is good for another few cycles of shrinkage, moving more into the third dimension by stacking increasing numbers of layers vertically, shrinking dimensions further by using extreme UV for lithography. How far can this take us? The technical problems of EUV are substantial, and have already absorbed major R&D investments. The current approaches for multiplying transistors will reach their end-point, whether killed by technical or economic problems, perhaps within the next decade.
Other physical substrates for computing are possible and are the subject of R&D at the moment, but none yet has a clear pathway for implementation. Quantum computing excites physicists, but we’re still some way from a manufacturable and useful device for general purpose computing.
There is one cause for optimism, though, which relates to energy consumption. There is a physical lower limit on how much energy it takes to carry out a computation – the Landauer limit. The plot above shows that our current technology for computing consumes energy at a rate which is many orders of magnitude greater than this theoretical limit (and for that matter, it is much more energy intensive than biological computing). There is huge room for improvement – the only question is whether we can deploy R&D resources to pursue this goal on the scale that’s gone into computing as we know it today.
See also Has Moore’s Law been repealed? An economist’s perspective, by Keith Flamm, in Computing in Science and Engineering, 2017

